Too Many False Targets for MicroRNAs: Challenges and Pitfalls in Prediction of miRNA Targets and Their Gene Ontology in Model and Non‐model Organisms - Fridrich - 2019 - BioEssays - Wiley Online Library
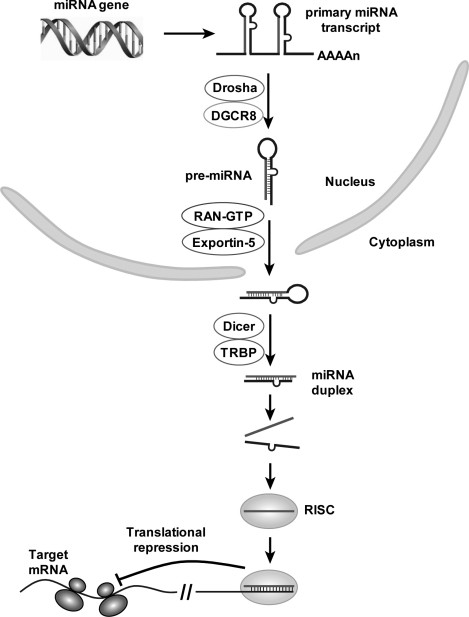
Got target?: computational methods for microRNA target prediction and their extension | Experimental & Molecular Medicine

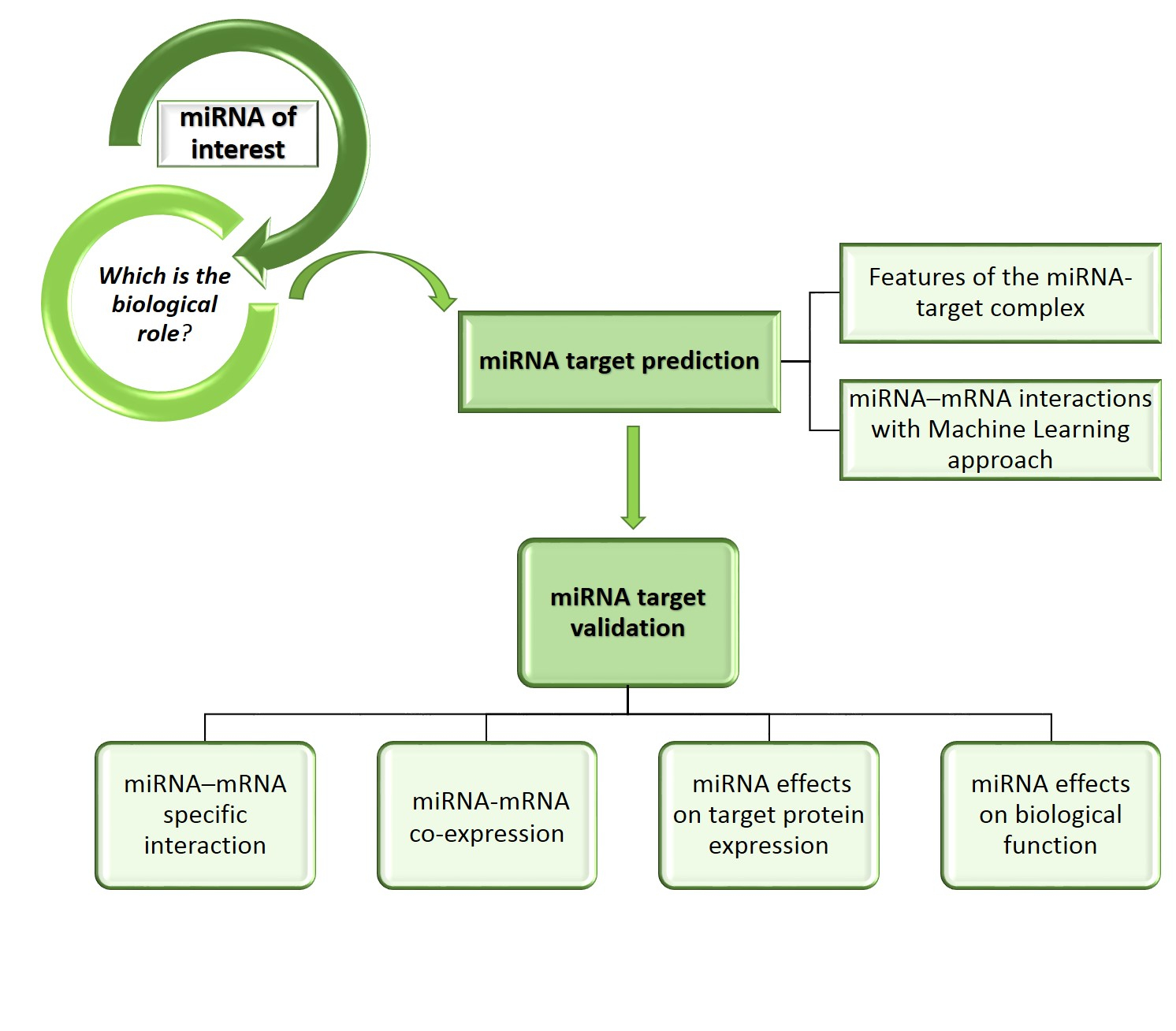
Target prediction and validation of microRNAs expressed from FSHR and aromatase genes in human ovarian granulosa cells | Scientific Reports

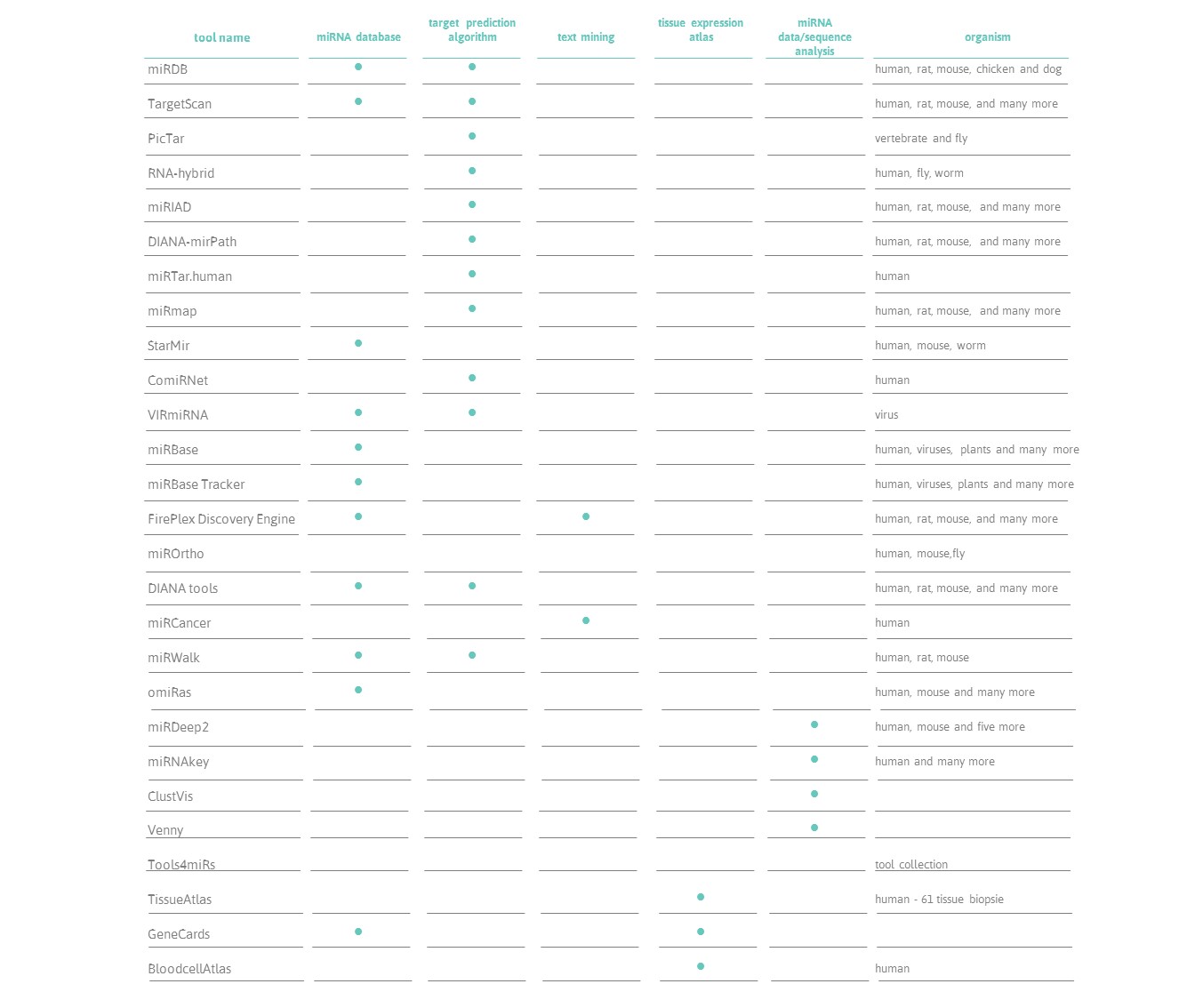
Popular Computational Tools Used for miRNA Prediction and Their Future Development Prospects | SpringerLink
![Computational identification of miRNAs that modulate the differentiation of mesenchymal stem cells to osteoblasts [PeerJ] Computational identification of miRNAs that modulate the differentiation of mesenchymal stem cells to osteoblasts [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2016/1976/1/fig-1-full.png)
Computational identification of miRNAs that modulate the differentiation of mesenchymal stem cells to osteoblasts [PeerJ]
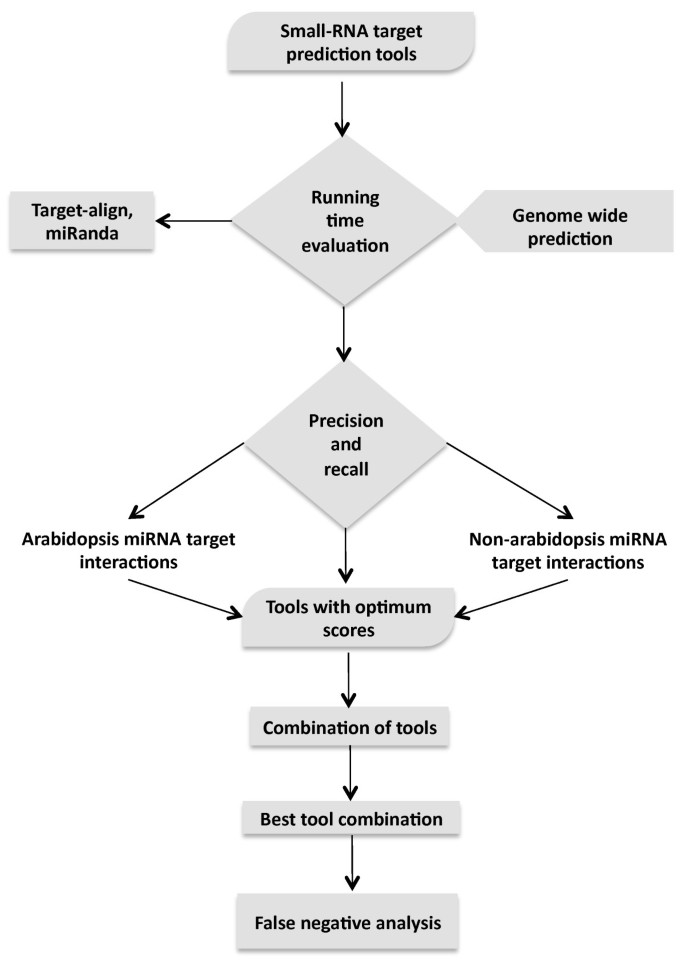
A comparison of performance of plant miRNA target prediction tools and the characterization of features for genome-wide target prediction | BMC Genomics | Full Text
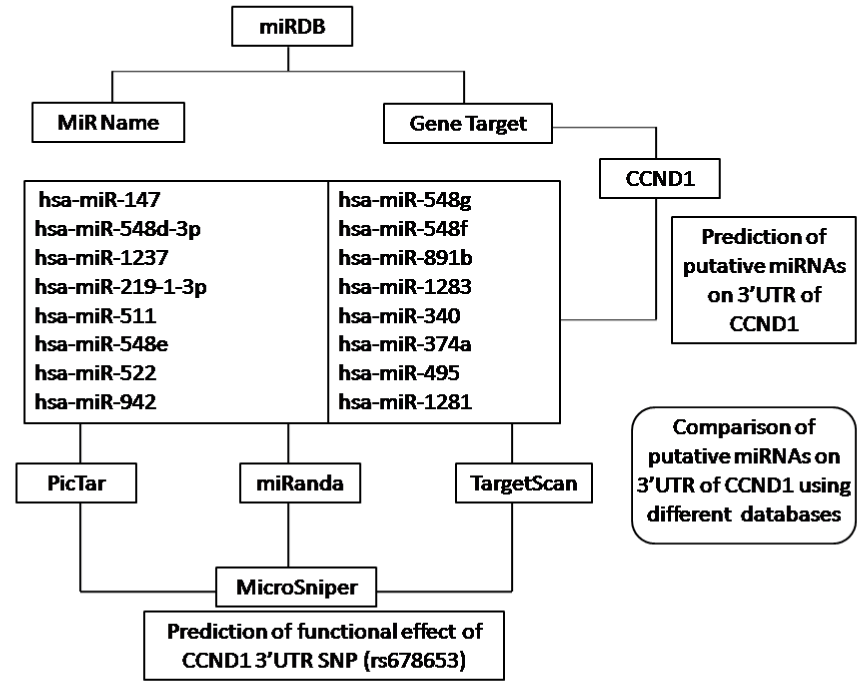
Computational prediction of novel human miRNAs/miRNA target sites in correlation with SNP (rs678653) at 3'UTR of cyclin D1 gene
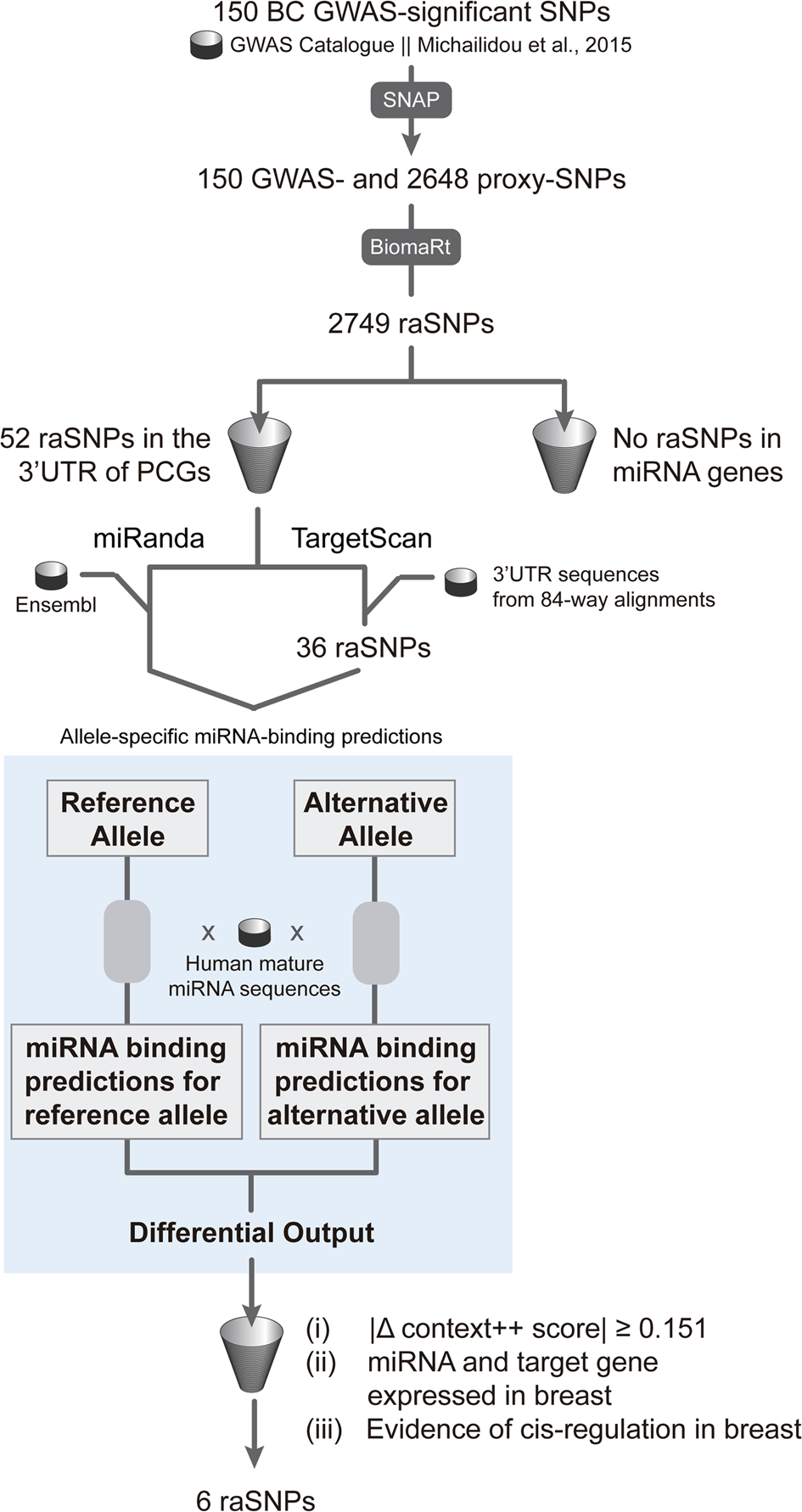
Allele-specific miRNA-binding analysis identifies candidate target genes for breast cancer risk | npj Genomic Medicine

MIPDH: A Novel Computational Model for Predicting microRNA–mRNA Interactions by DeepWalk on a Heterogeneous Network | ACS Omega

Prediction methods for microRNA targets in bilaterian animals: Toward a better understanding by biologists - ScienceDirect


![PDF] Computational Methods for MicroRNA Target Prediction | Semantic Scholar PDF] Computational Methods for MicroRNA Target Prediction | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6683bfaaec120a340fd0b2c4f3021d3691ea2029/5-Table1-1.png)




![PDF] Tools for Sequence-Based miRNA Target Prediction: What to Choose? | Semantic Scholar PDF] Tools for Sequence-Based miRNA Target Prediction: What to Choose? | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/35d57140ba819134565fb78da59485f5245a8413/6-Table1-1.png)